
Personal Information
Education: B.S. Molecular Biology, University of California, San Diego 2016
From: San Francisco, California
Joined the David Lab: March 2017
Outside of lab: Outside of lab, I enjoy going to the beach (while using proper sunscreen – beware of DNA damage!), eating, and pretending I can do photography.
Research Experience
Research in David Lab
My main project in the lab focuses on the interdomain connector (IDC) of mammalian homologs of the adenine glycosylase MUTYH. Mutations in the MUTYH gene are known to cause MUTYH associated polyposis (MAP) and contribute to an individual’s predisposition to colorectal cancer. These mutations are located throughout the protein, including the IDC. In the prokaryotic homologs, MutY, this region of the protein wraps around the DNA duplex upon MutY binding. The mammalian homologs have an extended interdomain linker, and this region is also known to be the docking site of various protein-protein interactions involving MUTYH. These higher eukaryotic homologs of MUTYH, including human and mouse, also possess a second metal cofactor in this IDC region, a Zn2+ ion coordinated by residues that comprise a Zn2+ linchpin motif. My project utilizes these MAP variants in the MUTYH IDC to understand what interactions are necessary for downstream DNA repair and signaling and how perturbations to this Zn2+ linchpin motif can alter those roles.
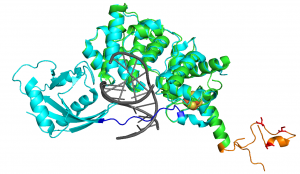
Alignment of partial human MUTYH structure with bacterial MutY. N-terminal fragment crystal structure of human MUTYH (PDB 3N5N, green) with IDC (orange) aligned to homolog structure of E. coli MutY (PDB 5DPK, light blue) bound to DNA (grey), highlighting the significantly shorter IDC found in prokaryotes (dark blue). Established Zn2+ ion chelating ligands Cys318, Cys325 and Cys328 are depicted in red. The Fe-S cluster is depicted as orange and yellow spheres. (Figure from Nuñez et al. JACS 2018)
Publications from David Lab
Previous Research Experience
My first research experience was at Sutro Biopharma, a biotechnology company in South San Francisco, where I first learned molecular biology techniques and fell in love with research. During undergrad, I was a UC LEADS scholar, which funded me to work in different labs during the school year and for two summers. At UCSD, I worked in Dr. Eva-Maria Collins’s lab, where I helped coordinate data collection and analysis of a regeneration assay to study planarian regeneration in the presence of different toxins and toxicants. During summer 2015, I worked with Dr. Jorge Torres at UCLA to look at how an E3 ubiquitin ligase could be involved in mitosis and cancer progression.
Publications from Previous Research
Updated: April 13, 2021
